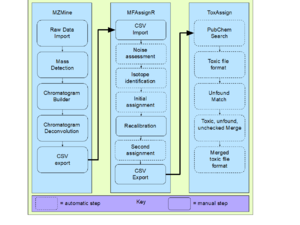
Previous published methods for non-targeted screening of toxins in alternative foods such as leaf concentrate, agricultural residues or plastic fed to biological consortia are time consuming and expensive and thus present accessibility, as well as, time-constraint issues for scientists from under resourced settings to identify safe alternative foods. The novel methodology presented here, utilizes a completely free and open source software toolchain for automatically screening unknown alternative foods for toxicity using experimental data from high-pressure liquid chromatography and mass spectrometry. The process uses three distinct tools (mass spectrometry analysis with MZmine 2, formula assignment with MFAssignR, and data filtering with ToxAssign) enabling it to be modular and easily upgradable in the future. MZmine 2 and MFAssignR have been previously described, while ToxAssign was developed here to match the formulas output by formula assignment to potentially toxic compounds in a local table, then look up toxic data on the Open Food Tox Database for the matched compounds. This process is designed to fill the gap between food safety analysis techniques and developing alternative food production techniques to allow for new methods of food production to be preliminarily tested before animal testing. The methodology was validated against a previous method using proprietary commercial software. The new process identifies all of the toxic elements the previous process identified with more detailed information than the previous process was able to provide automatically.
- Efficient analysis to find potentially toxic compounds in alternative foods
- Identification of potentially unsafe products without the use of live animal testing
- Modular design to allow for upgrading or fitting of user needs
- Source code: https://osf.io/nh76z/
- MZmine 2 https://mzmine.github.io/
- MFAssignR https://github.com/skschum/MFAssignR
- ToxAssign https://osf.io/nh76z/
See also[edit | edit source]
- Feeding Everyone No Matter What - The full book main page
- David Denkenberger and Joshua Pearce, Feeding Everyone No Matter What: Managing Food Security After Global Catastrophe , 1st Edition, Academic Press, 2015
- Free Preview: Google books
- Cover on Academia
- Facebook page
- Alternative Foods as a Solution to Global Food Supply Catastrophes
- Resilience to global food supply catastrophes
- Feeding Everyone if the Sun is Obscured and Industry is Disabled
- Cost-Effectiveness of Interventions for Alternate Food to Address Agricultural Catastrophes Globally
- Feeding Everyone: Solving the Food Crisis in Event of Global Catastrophes that Kill Crops or Obscure the Sun
- Food without sun: Price and life-saving potential
- Cost-effectiveness of interventions for alternate food in the United States to address agricultural catastrophes
- Micronutrient Availability in Alternative Foods During Agricultural Catastrophes
- Preliminary Automated Determination of Edibility of Alternative Foods: Non-Targeted Screening for Toxins in Red Maple Leaf Concentrate
- Open Source Software Toolchain for Automated Non-Targeted Screening for Toxins in Alternative Foods
- Scaling of greenhouse crop production in low sunlight scenarios
- Potential of microbial protein from hydrogen for preventing mass starvation in catastrophic scenarios
- U.S. Potential of Sustainable Backyard Distributed Animal and Plant Protein Production During & After Pandemics
- Global distribution of forest classes and leaf biomass for use as alternative foods to minimize malnutrition
- Long-term cost-effectiveness of interventions for loss of electricity/industry compared to artificial general intelligence safety
- Long term cost-effectiveness of resilient foods for global catastrophes compared to artificial general intelligence safety
- Rapid repurposing of pulp and paper mills, biorefineries, and breweries for lignocellulosic sugar production in global food catastrophes
- Nutrition in Abrupt Sunlight Reduction Scenarios: Envisioning Feasible Balanced Diets on Resilient Foods
- Methane Single Cell Protein: securing protein supply during global food catastrophes
- Killing two birds with one stone: chemical and biological upcycling of polyethylene terephthalate plastics into food
- How Easy is it to Feed Everyone? Economic Alternatives to Eliminate Human Nutrition Deficits
- Quantifying Alternative Food Potential of Agricultural Residue in Rural Communities of Sub-Saharan Africa
- Yield and Toxin Analysis of Leaf Protein Concentrate from Common North American Coniferous Trees
- Toxic Analysis of Leaf Protein Concentrate Regarding Common Agricultural Residues
- Towards Sustainable Protein Sources: The Thermal and Rheological Properties of Alternative Proteins
Additional Information[edit source]
- ALLFED
- Dave Denkenberger Publications
- OSE Wiki "Synfood" (i.e. protein and other dietary components from microbial organisms fed on gas or other hydrocarbons)
Davos IDRC Conference[edit source]
- Papers
- MOST completed projects and publications
- Famine relief
- Farming
- Food
- Food and agriculture
- Food crops
- Food processing
- Food production
- Food safety
- Food security
- Food storage
- Resilient foods
- SDG02 Zero hunger
- SDG12 Responsible consumption and production
- Open source software
- FAST Completed
- Sustainable development
- Public health





